Environmental DNA Is Everywhere. Scientists Are Gathering It All.
In the late 1980s, at a federal research facility in Pensacola, Florida, Tamar Barkay used mud in a way that proved revolutionary in a manner she could never have imagined at the time: a crude version of a technique that is now shaking up many scientific fields. Barkay had collected several samples of mud — one from an inland reservoir, another from a brackish bayou, and a third from a low-lying saltwater swamp. She put these sediment samples in glass bottles in the lab, and then added mercury, creating what amounted to toxic sludge.

At the time, Barkay worked for the Environmental Protection Agency and she wanted to know how microorganisms in mud interact with mercury, an industrial pollutant, which required an understanding of all the organisms in a given environment — not just the tiny portion that could be successfully grown in Petri dishes in the lab. But the underlying question was so basic that it remains one of those fundamental driving queries across biology. As Barkay, who is now retired, put it in a recent interview from Boulder, Colorado: “Who is there?” And, just as important, she added: “What are they doing there?”
Such questions are still relevant today, asked by ecologists, public health officials, conservation biologists, forensic practitioners, and those studying evolution and ancient environments — and they drive shoe-leather epidemiologists and biologists to far-flung corners of the world.
The 1987 paper Barkay and her colleagues published in the Journal of Microbiological Methods outlined a method — “Direct Environmental DNA Extraction” — that would allow researchers to take a census. It was a practical tool, albeit a rather messy one, for detecting who was out there. Barkay used it for the rest of her career.
Today, the study gets cited as an early glimpse of eDNA, or environmental DNA, a relatively inexpensive, widespread, potentially automated way to observe the diversity and distribution of life. Unlike previous techniques, which could identify DNA from, say, a single organism, the method also collects the swirling cloud of other genetic material that surrounds it. In recent years, the field has grown significantly. “It’s got its own journal,” said Eske Willerslev, an evolutionary geneticist at the University of Copenhagen. “It’s got its own society, scientific society. It has become an established field.”
“We’re all flaky, right? There’s bits of cellular debris sloughing off all the time.”
eDNA serves as a surveillance tool, offering researchers a means of detecting the seemingly undetectable. By sampling eDNA, or mixtures of genetic material — that is, fragments of DNA, the blueprint of life — in water, soil, ice cores, cotton swabs, or practically any environment imaginable, even thin air, it is now possible to search for a specific organism or assemble a snapshot of all the organisms in a given place. Instead of setting up a camera to see who crosses the beach at night, eDNA pulls that information out of footprints in the sand. “We’re all flaky, right?” said Robert Hanner, a biologist at the University of Guelph in Canada. “There’s bits of cellular debris sloughing off all the time.”
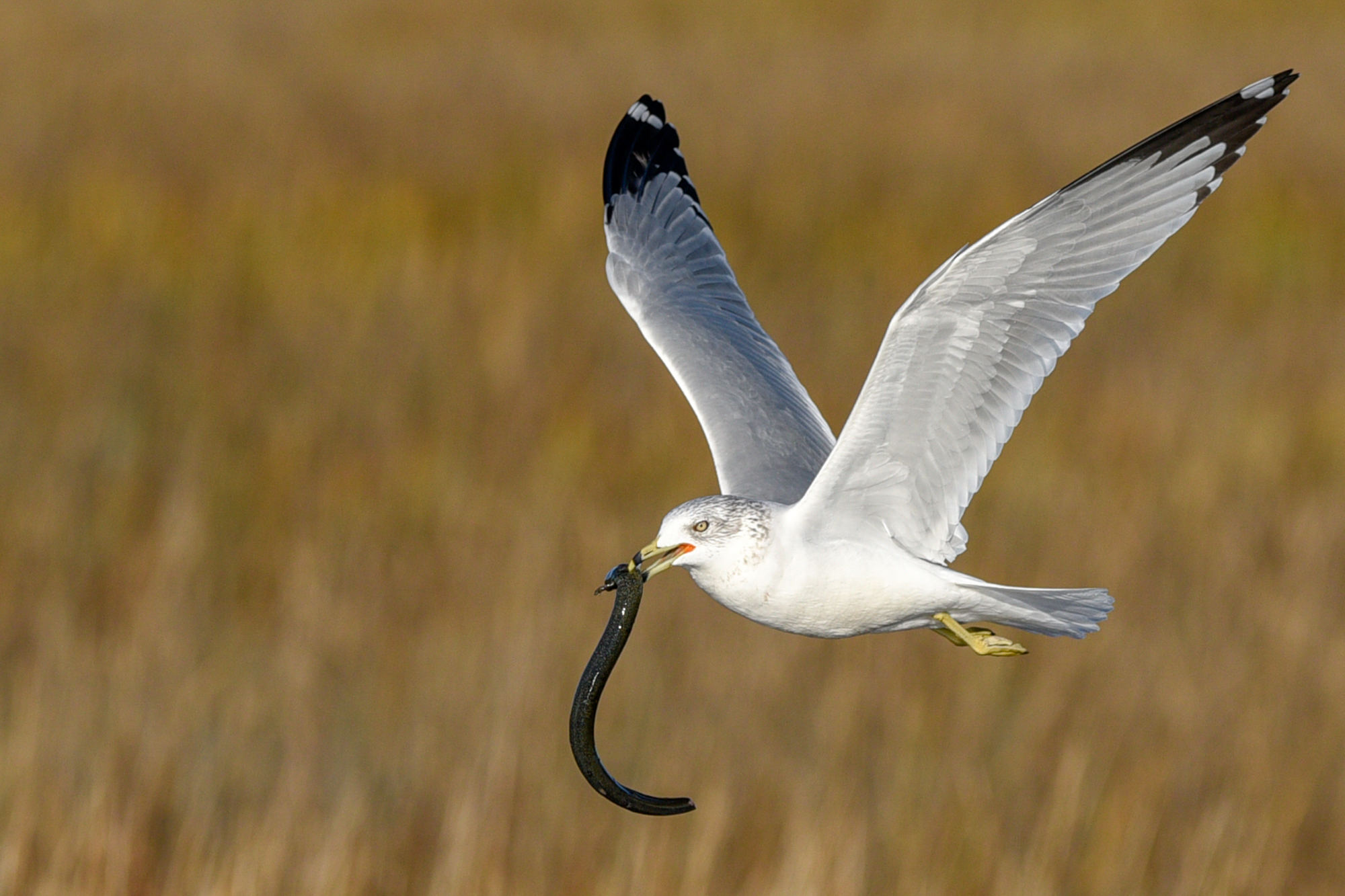
As a method for confirming the presence of something, eDNA isn’t failproof. For instance, the organism detected in eDNA might not actually live in the location where the sample was collected; Hanner gave the example of a passing bird, a heron, that ate a salamander and then pooped out some of its DNA, which could be one reason signals of the amphibian are present in some areas where they’ve never been physically found.
Still, eDNA has the ability to help sleuth out genetic traces, some of which slough off in the environment, offering a thrilling — and potentially chilling — way to collect information about organisms, including humans, as they go about their everyday business.
The conceptual basis for eDNA — pronounced EE-DEE-EN-AY, not ED-NUH — dates back a hundred years, before the advent of so-called molecular biology, and it is often attributed to Edmond Locard, a French criminologist working in the early 20th century. In a series of papers published in 1929, Locard proposed a principle: Every contact leaves a trace. In essence, eDNA brings Locard’s principle to the 21st century.
For the first several decades, the field that became eDNA — Barkay’s work in the 1980s included — focused largely on microbial life. Looking back at its evolution, eDNA appeared slow to claw its way out of the proverbial mud.

It wasn’t until 2003 that the method turned up a vanished ecosystem. Led by Willerslev, the 2003 study pulled ancient DNA from less than a teaspoon of sediment, demonstrating for the first time the feasibility of detecting larger organisms with the technique, including plants and woolly mammoths. In the same study, sediment collected in a New Zealand cave (which notably had not been frozen) revealed an extinct bird: the moa. What is perhaps most remarkable is that these applications for studying ancient DNA stemmed from a prodigious amount of dung dropped on the ground hundreds of thousands of years ago.
Willerslev had first come up with the idea a few years earlier while contemplating a more recent pile of dung: In between his master’s degree and Ph.D. in Copenhagen, he found himself at loose ends, struggling to obtain bones, skeletal remains, or other physical specimens to study. But one autumn, he gazed out the window at “a dog taking a crap on the street,” he recalled. The scene prompted him to think about the DNA in feces, and how it washed away with rain, leaving no visible trace. But Willerslev wondered, “‘Could it be that the DNA could survive?’ That’s what I then set up to try to find out.”
The paper demonstrated the remarkable persistence of DNA, which, he said, survives in the environment for much longer than previous estimates suggested. Willerslev has since analyzed eDNA in frozen tundra in modern-day Greenland, dating back 2 million years ago, and he is working on samples from Angkor Wat, the enormous temple complex in Cambodia believed to have been built in the 12th century. “It should be the worst DNA preservation you can imagine,” he said. “I mean, it’s hot and humid.”
But, he said, “we can get DNA out.”
eDNA has the ability to help sleuth out genetic traces, offering a thrilling — and potentially chilling — way to collect information about organisms as they go about their everyday business.
Willerslev is now hardly alone in seeing a potential tool with seemingly limitless applications — especially now as advances enable researchers to sequence and analyze larger quantities of genetic information. “It’s an open window for many, many things,” he said, “and much more than I can think of, I’m sure.” It was not just ancient mammoths; eDNA could reveal present-day organisms hiding in our midst.
Scientists use eDNA to track creatures of all shapes and sizes, be it a single species, such as tiny bits of invasive algae, eels in Loch Ness, or a sightless sand-dwelling mole that hasn’t been seen in nearly 90 years; researchers sample entire communities, say, by looking at the eDNA found on wildflower blossoms or the eDNA blowing in the wind as a proxy for all the visiting birds and bees and other animal pollinators.
The next evolutionary leap forward in eDNA’s history took shape around the search for organisms currently living in earth’s aquatic environments. In 2008, a headline appeared: “Water retains DNA memory of hidden species.” It came not from the supermarket tabloid, but the respected trade publication Chemistry World, describing work by French researcher Pierre Taberlet and his colleagues. The group sought out brown-and-green bullfrogs, which can weigh more than 2 pounds and, because they mow down everything in their path, are considered an invasive species in western Europe. Finding bullfrogs usually involved skilled herpetologists scanning shorelines with binoculars who then returned after sunset to listen for their calls. The 2008 paper suggested an easier way — a survey that required a lot less personnel.
“You could get DNA from that species directly out of the water,” said Philip Thomsen, a biologist at Aarhus University (who was not involved in the study). “And that really kickstarted the field of environmental DNA.”
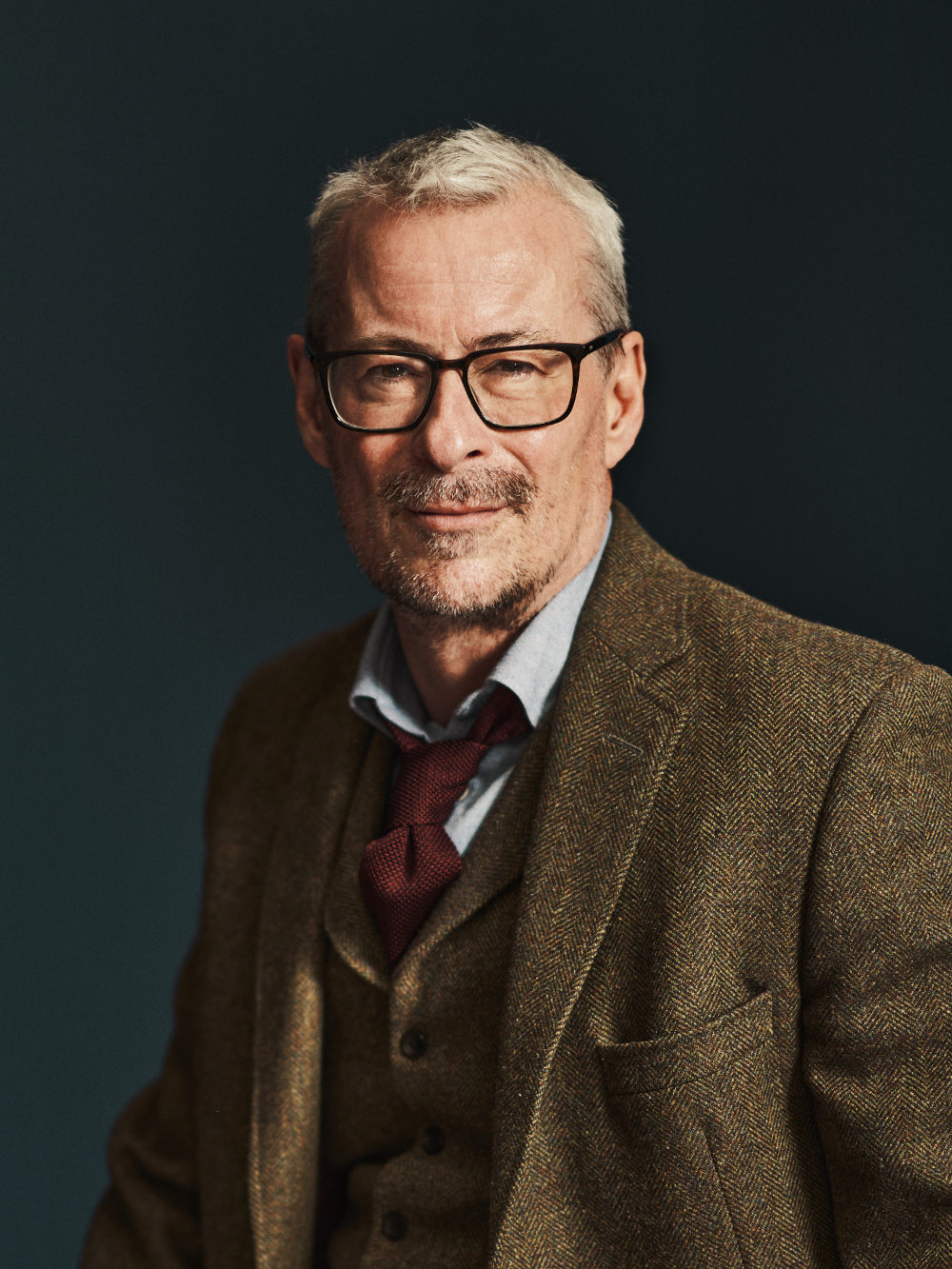
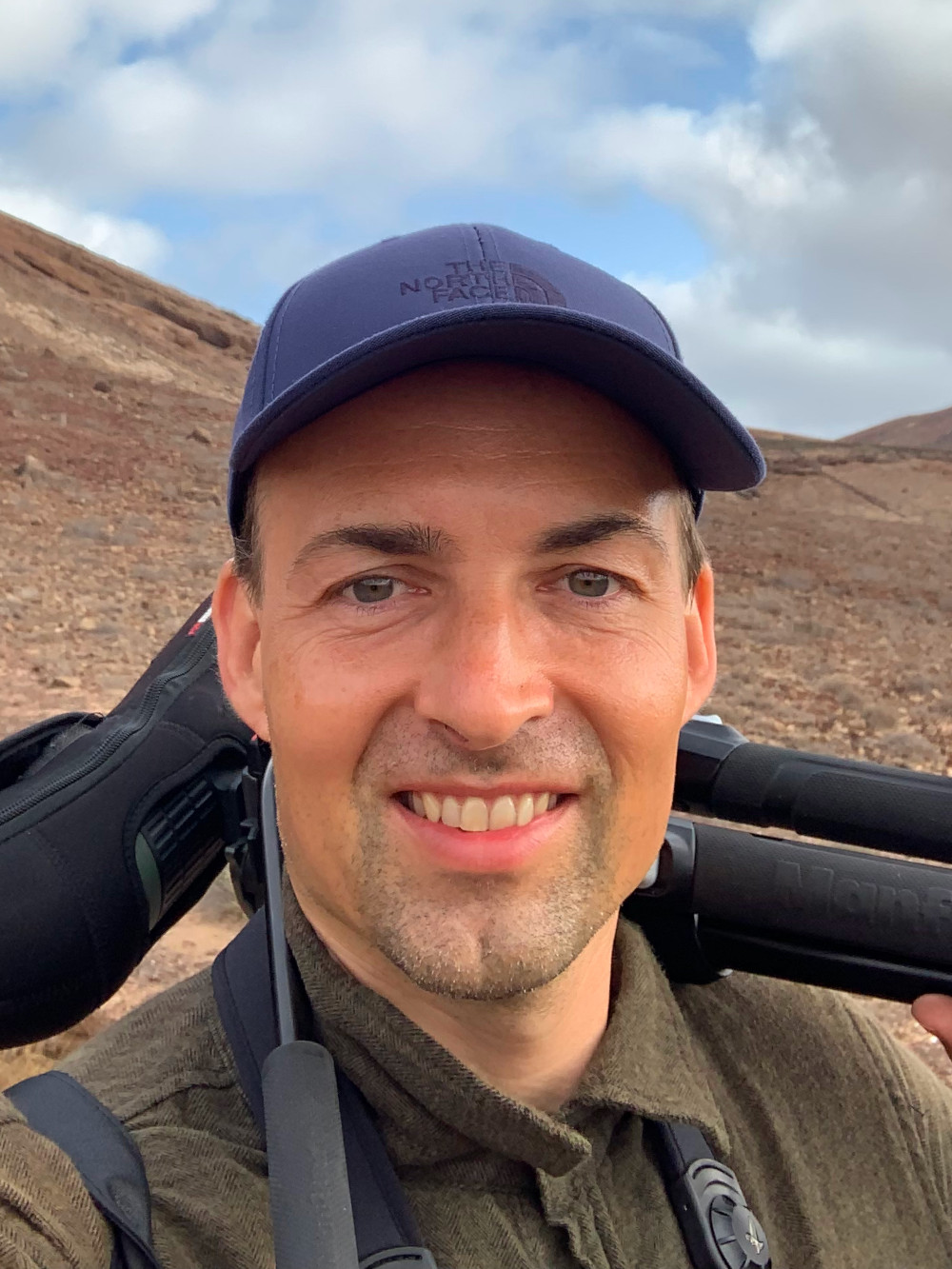



Frogs can be hard to detect, and they are not, of course, the only species that eludes more traditional, boots-on-the-ground detection. Thomsen began work on another organism that notoriously confounds measurement: fish. Counting fish is sometimes said to vaguely resemble counting trees — except they’re free-roaming, in dark places, and fish counters are doing their tally while blindfolded. Environmental DNA dropped the blindfold. One review of published literature on the technology — though it came with caveats, including imperfect and imprecise detections or details on abundance — found that eDNA studies on freshwater and marine fish and amphibians outnumbered terrestrial counterparts 7:1.
In 2011, Thomsen, then a Ph.D. candidate in Willerslev’s lab, published a paper demonstrating that the method could detect rare and threatened species, such as those in low abundance in Europe, including amphibians, mammals like the otter, crustaceans, and dragonflies. “We showed that only, like, a shot glass of water really was enough to detect these organisms,” he told Undark. It was clear: The method had direct applications in conservation biology for the detection and monitoring of species.
In 2012, the journal Molecular Ecology published a special issue on eDNA, and Taberlet and several colleagues outlined a working definition of eDNA as any DNA isolated from environmental samples. The method described two similar but slightly different approaches: One can answer a yes or no question: Is the bullfrog (or whatever) present or not? It does so by scanning the metaphoric barcode, short sequences of DNA that are particular to a species or family, called primers; the checkout scanner is a common technique called quantitative real-time polymerase chain reaction, or qPCR.
Scientists use eDNA to track creatures of all shapes and sizes, be it tiny bits of invasive algae, eels in Loch Ness, or a sightless sand-dwelling mole that hasn’t been seen in nearly 90 years.
Another approach, commonly known as DNA metabarcoding, essentially spits out a list of organisms present in a given sample. “You sort of ask the question, what is here?” Thomsen said. “And then you get all of the known things, but you also get some surprises, right? Because there were some species that you didn’t know were actually present.”
One aims to find the needle in a haystack; the other attempts to reveal the whole haystack. eDNA differs from more traditional sampling techniques where organisms, like fish, are caught, manipulated, stressed, and sometimes killed. The data obtained are objective; it’s standardized and unbiased.
“eDNA, one way or the other, is going to stay as one of the important methodologies in biological sciences,” said Mehrdad Hajibabaei, a molecular biologist at University of Guelph, who pioneered the metabarcoding approach, and who traced fish some 9,800 feet under the Labrador Sea. “Every day I see something bubbling up that didn’t occur to me.”
In recent years, the field of eDNA has expanded. The method’s sensitivity allows researchers to sample previously out-of-reach environments, for example, capturing eDNA from the air — an approach that highlights eDNA’s promises and its potential pitfalls. Airborne eDNA appears to circulate on a global dust belt, suggesting its abundance and omnipresence, and it can be filtered and analyzed to monitor plants and terrestrial animals. But eDNA blowing in the wind can lead to inadvertent contamination.
In 2019, Thomsen, for instance, left two bottles of ultra-pure water out in the open — one in a grassland, and the other near a marine harbor. After a few hours, the water contained detectable eDNA associated with birds and herring, suggesting that traces of non-terrestrial species settled into the samples; the organisms obviously did not inhabit the bottles. “So it must come from the air,” Thomsen told Undark. The results suggest a two-fold problem: For one, trace evidence can move around, where two organisms that come into contact can then tote around the other’s DNA, and just because certain DNA is present doesn’t mean that the species is actually there.
Moreover, there’s also no guarantee that the presence of eDNA indicates that a species is alive, and field surveys are still needed, he said, to understand a species’ breeding success, its health, or the status of its habitat. So far, then, eDNA does not necessarily replace physical observations or collections. In another study, in which Thomsen’s group collected eDNA on flowers to look for pollinating birds, more than half of the eDNA reported in the paper came from humans, contamination that potentially muddied the results and made it harder to detect the pollinators in question.
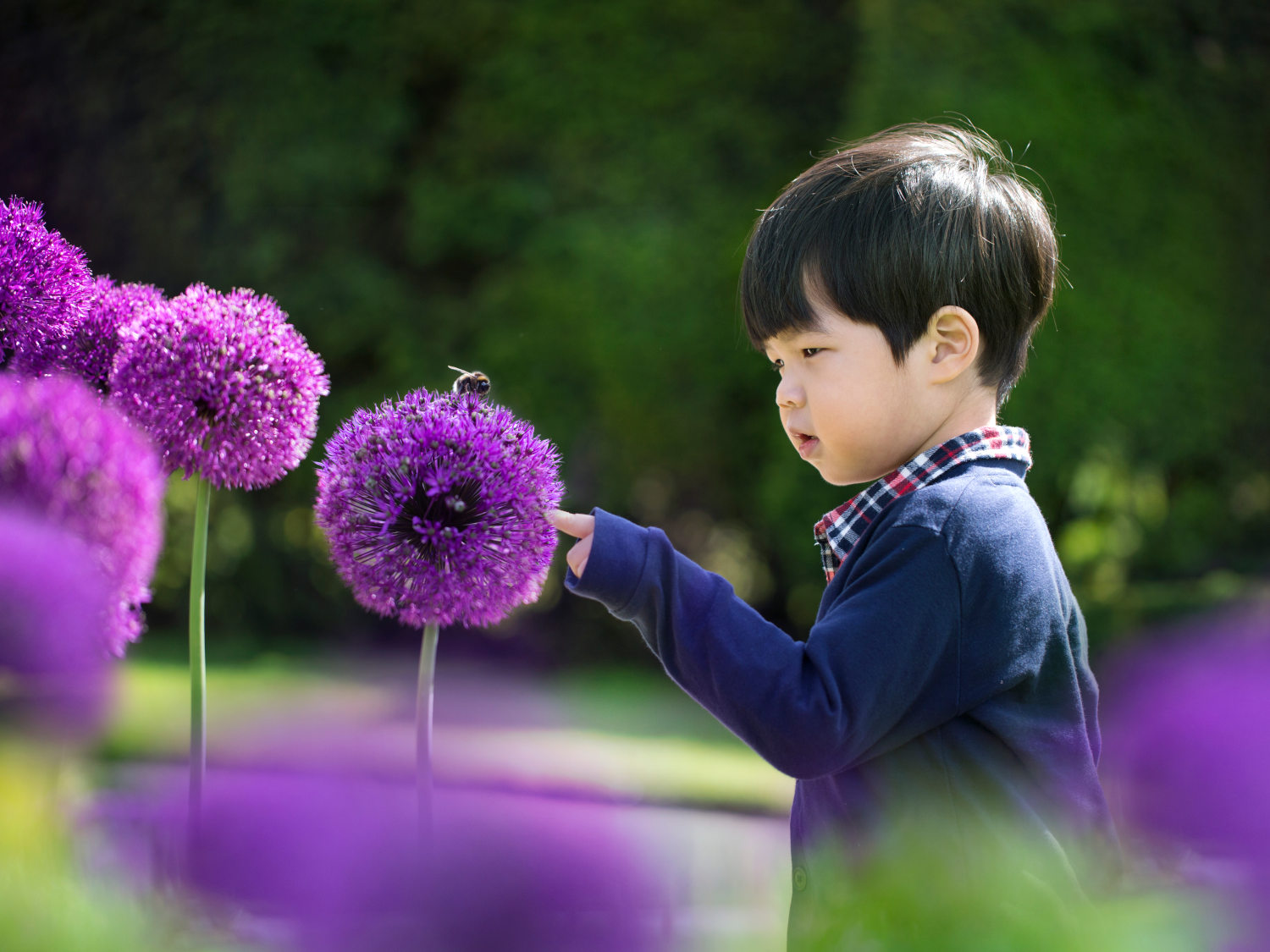
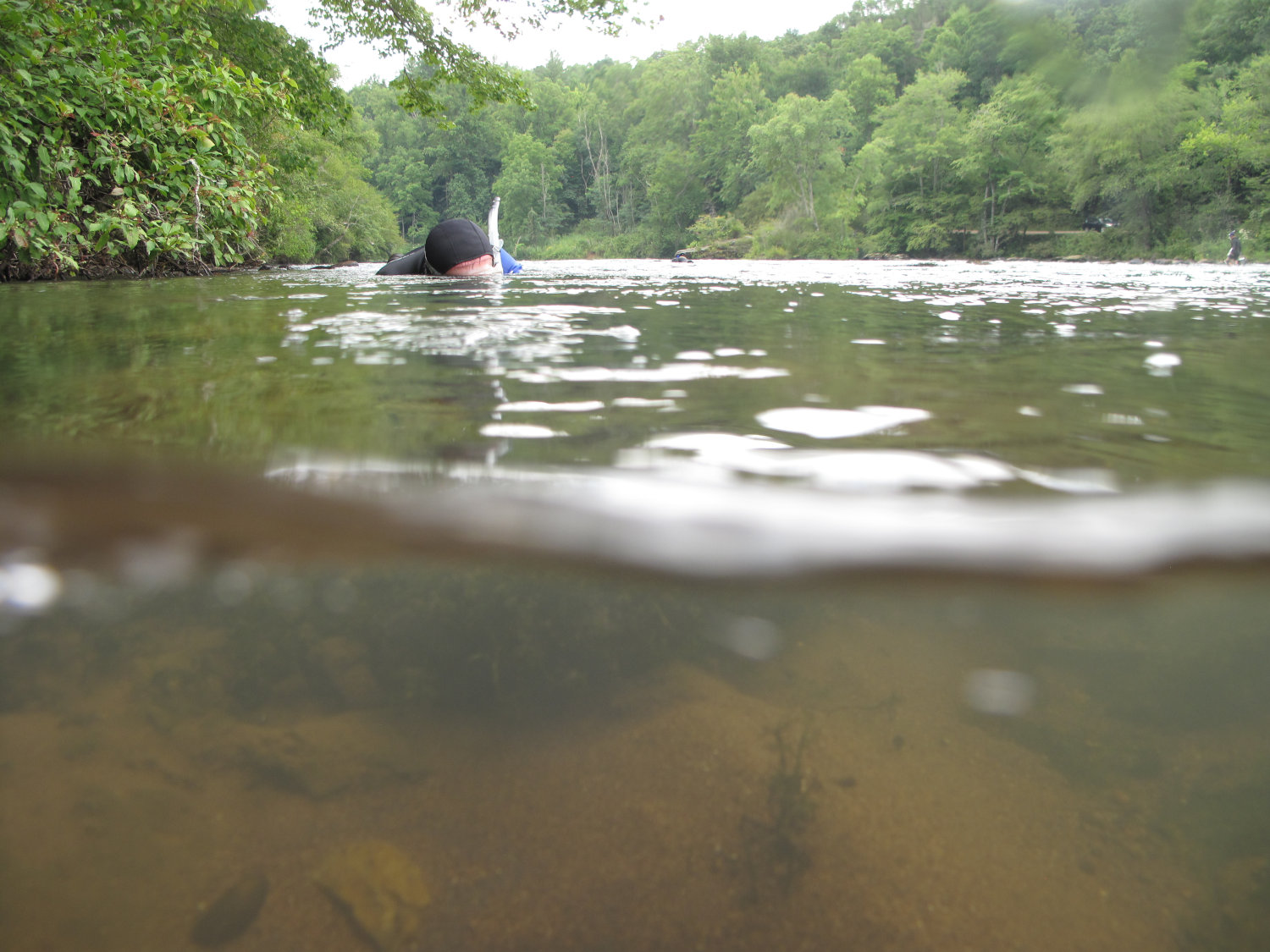


Similarly, in May 2023, a University of Florida team that previously studied sea turtles by the eDNA traces left as they crawl along the beach published a paper that turned up human DNA. The samples were intact enough to detect key mutations that might someday be used to identify individual people, suggesting that the biological surveillance also raised unanswered questions about ethical testing on humans and informed consent. If eDNA served as a seine net, then it indiscriminately swept up information about biodiversity and inevitably ended up with, as the UF team’s paper put it, “human genetic by-catch.”
While the privacy issues around footprints in the sand, so far, appear to exist mostly in the realm of hypothetical, the use of eDNA in legal litigation relating to wildlife is not only possible but already a reality. It’s also being used in criminal investigations: In 2021, for instance, a group of Chinese researchers reported that eDNA collected off a suspected murderer’s pants had, contrary to his claims, revealed that he’d likely been to the muddy canal where a dead body had been found.
The concerns about off-target eDNA, in terms of accuracy and its reach into human medicine and forensics, highlight another, much broader, shortcoming. As Hanner at the University of Guelph described the problem: “Our regulatory frameworks and policy tend to lag at least a decade or more behind the science.”
“Every day I see something bubbling up that didn’t occur to me.”
Today, there are countless potential regulatory applications for water quality monitoring, evaluating environmental impact (including offshore wind farms and oil and gas drilling to more run-of-the-mill strip mall development), species management, and enforcement of the Endangered Species Act. In a civil court case filed in 2021, the U.S. Fish and Wildlife Service evaluated whether an imperiled fish existed in a particular watershed, using eDNA and more traditional sampling, and found that they did not. The courts said the agency’s lack of protections for that watershed were justified. The issue does not seem to be whether eDNA stood up in court; it did. “But you really can’t say that something does not exist in an environment,” said Hajibabaei.
He recently highlighted the issue of validation: eDNA infers a result, but needs more established criteria for confirming that these results are actually true (that an organism is actually present or absent, or in a certain quantity). A series of special meetings for scientists worked to address these issues of standardization, which he said include protocols, chain of custody, and criteria for data generation and analysis. In a review of eDNA studies, Hajibabaei and his colleagues found that the field is saturated with one-offs, or proof-of-concept studies attempting to show that eDNA analyses work. Research remains overwhelmingly siloed in academia.
As such, practitioners hoping to use eDNA in an applied contexts sometimes ask for the moon. Does the species exist in certain location? For instance, Hajibabaei said, someone recently asked him if he could totally refute the presence of a parasite, proving that it had not appeared in an aquaculture farm. “And I say, ‘Look, there is no way that I can say that is 100 percent.’”



Even with a rigorous analytic framework, he said, the issues with false negatives and false positives are particularly difficult to resolve without doing one of the things eDNA obviates — more traditional collection and manual inspection. Despite the limitations, a handful of companies are already starting to commercialize the technique. For instance, future applications could help a company confirm whether the bridge it is building will harm any locally endangered animals; an aquaculture outfit determine if the waters where it farms its fish are infested with sea lice; or a landowner who is curious whether new plantings are attracting a wider range of native bees.
The problem is rather fundamental given eDNA’s reputation as an indirect way of detecting the undetectable — or as a workaround in contexts when it’s simply not possible to dip a net and catch all the organisms in the sea.
“It is very hard to validate some of these scenarios,” Hajibabaei said. “And that’s basically the nature of the beast.”
eDNA opens up a lot of possibilities, answering a question originally posed by Barkay (and no doubt many others): “Who is there?” But increasingly it’s providing hints that get at the “What are they doing there?” question, too. Elizabeth Clare, a professor of biology at York University in Toronto, studies biodiversity. She said she has observed bats roosting in one spot during the day, but, by collecting airborne eDNA, she could also infer where bats socialize at night. In another study, domesticated dog eDNA turned up in red fox scat. The two canids did not appear to be interbreeding, but researchers did wonder if their closeness had led to confusion, or cross-contamination, before ultimately settling on another explanation: Foxes apparently ate dog poop.
So while eDNA does not inherently reveal animal behavior, by some accounts the field is making strides towards providing clues as to what an organism might be doing, and how it’s interacting with other species, in a given environment — gleaning information about health without directly observing behavior.
Support Undark Magazine
Undark is a non-profit, editorially independent magazine covering the complicated and often fractious intersection of science and society. If you would like to help support our journalism, please consider making a donation. All proceeds go directly to Undark’s editorial fund.
Take another possibility: large-scale biomonitoring. Indeed, for the last three years, more people than ever before have participated in a bold experiment that is already up and running: the collection of environmental samples from public sewers to track viral Covid-19 particles and other organisms that infect humans. Technically, wastewater sampling involves a related approach called eRNA, because some viruses only have genetic information stored in the form of RNA, rather than DNA. Still, the same principles apply. (Studies also suggest RNA, which determines which proteins an organism is expressing, could be used to assess ecosystem health; organisms that are healthy may express entirely different proteins compared to those that are stressed.) In addition to monitoring the prevalence of diseases, wastewater surveillance demonstrates how an existing infrastructure designed to do one thing — sewers were designed to collect waste — could be fashioned into a powerful tool for studying something else, like detecting pathogens.
Clare has a habit of doing just that. “I personally am one of those people who tends to use tools — not the way they were intended,” she said. Clare was among the researchers who noticed a gap in the research: There was a lot less eDNA work done on terrestrial organisms. So, she began working with what might be called a natural filter, that is worms that suck blood from mammals. “It’s a lot easier to collect 1,000 leeches than it is to find the animals. But they have blood-meals inside them and the blood carries the DNA of the animals they interacted with,” she said. “It’s like having a bunch of field assistants out surveying for you.” Then, one of her students thought the same thing for dung beetles, which are even easier to collect.
Clare is now spearheading a new application for another continuous monitoring system — leveraging existing air-quality monitors that measure pollutants, such as fine particulate matter, while also simultaneously vacuuming eDNA out of the sky. In late 2023, she only had a small sample set, but had already found that, as a byproduct of routine air quality monitoring, these preexisting tools doubled as filters for the material she is after. It was, more or less, a regulated, transcontinental network collecting samples in a very consistent way over long periods of time. “You could then use it to build up time series and high-resolution data on entire continents,” she said.
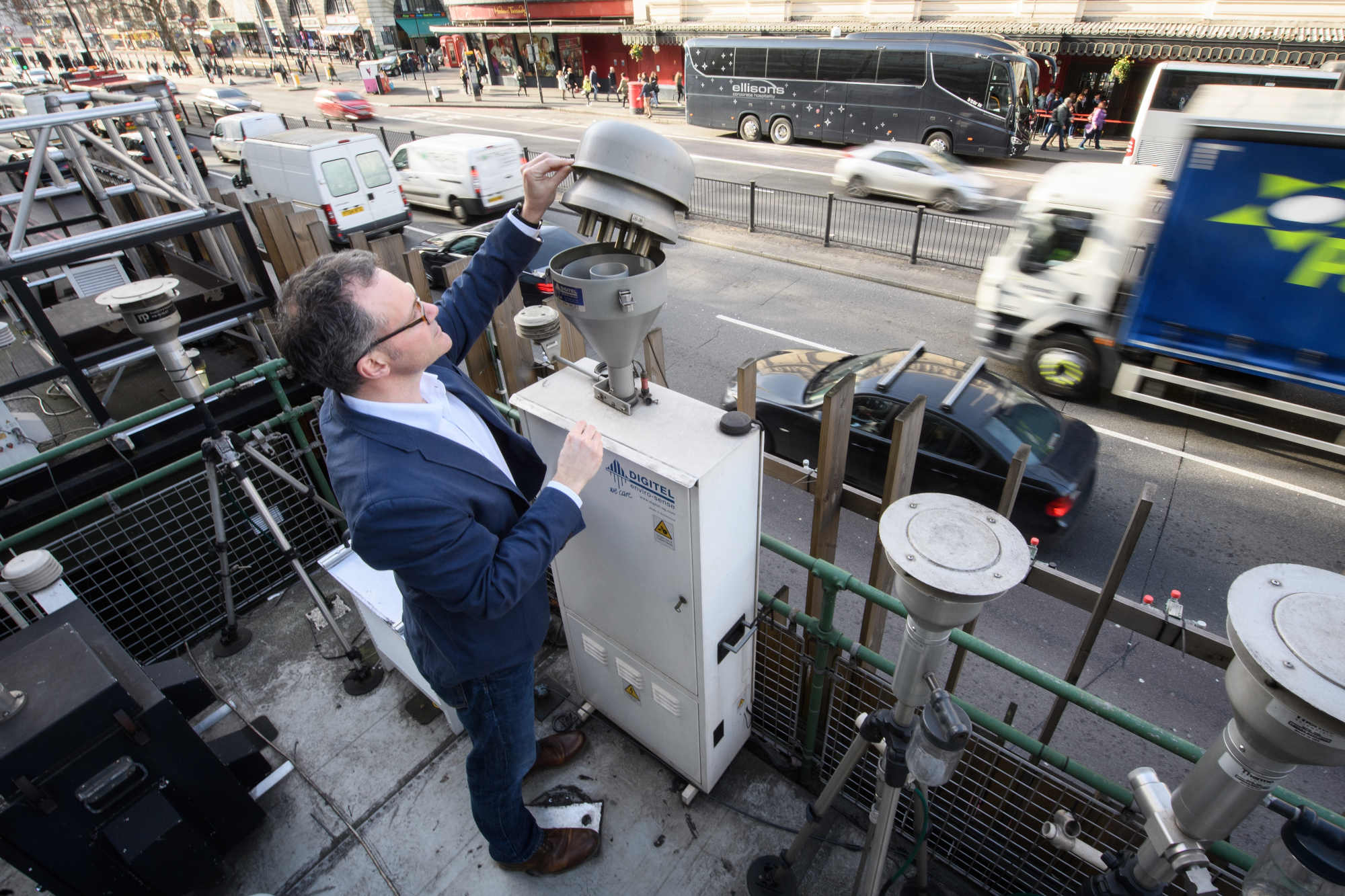

In the U.K. alone, Clare said, there are an estimated 150 different sites sucking a known quantity of air, every week, all year long, which amount to some 8,000 measurements a year. Clare and her co-authors recently analyzed at a tiny subset of these — 17 measurements from two locations — and were able to identify more than 180 different taxonomic groups, more than 80 different kinds of plants and fungi, 26 different species of mammal, 34 different species of birds, plus at least 35 kinds of insects.
Certainly, other long-term ecological research sites exist. The U.S. has a network of such facilities. But their scope of study does not include a globally distributed infrastructure that measures biodiversity constantly — including the passage of migrating birds overhead to the expansion and contraction of species with climate change. Arguably, eDNA will likely complement, rather than supplant, the distributed network of people, who record real-time, high-resolution, tempo-spatial observations on websites such as eBird or iNaturalist. Like a fuzzy image of an entirely new galaxy coming into view, the current resolution remains low.
“It’s sort of a generalized collection system, which is pretty much unheard of in biodiversity science,” said Clare. She was referring to the capacity to pull eDNA signals out of thin air, but the sentiment spoke to the method as a whole: “It’s not perfect,” she said, “but there’s nothing else that really does that.”
SHREDS OF EVIDENCE: THE COMPLETE SERIES
Part 1: DNA Deluge
Part 2: Hidden Bugs
Part 3: Evasive Traces
Part 4: Market Buzz
Part 5: Genetic Net
Part 6: Lost Souls